BACTERIOPHAGES Versus EUKARYOTIC Viruses (like COVID/Spike) AND WHICH KIND Infects What: (Spoiler Alert--the Bifidobacterium are most likely safe from COVID/Spike Protein)
When I was just a kid, long ago, the very first thing I learned that led me to become fascinated with science was learning about bacteriophages (also known as phages).
The bacteriophage looked like a mosquito to me—I was amazed at what it did—it also looked a bit scary to me as well—like something out of a sci fi movie.
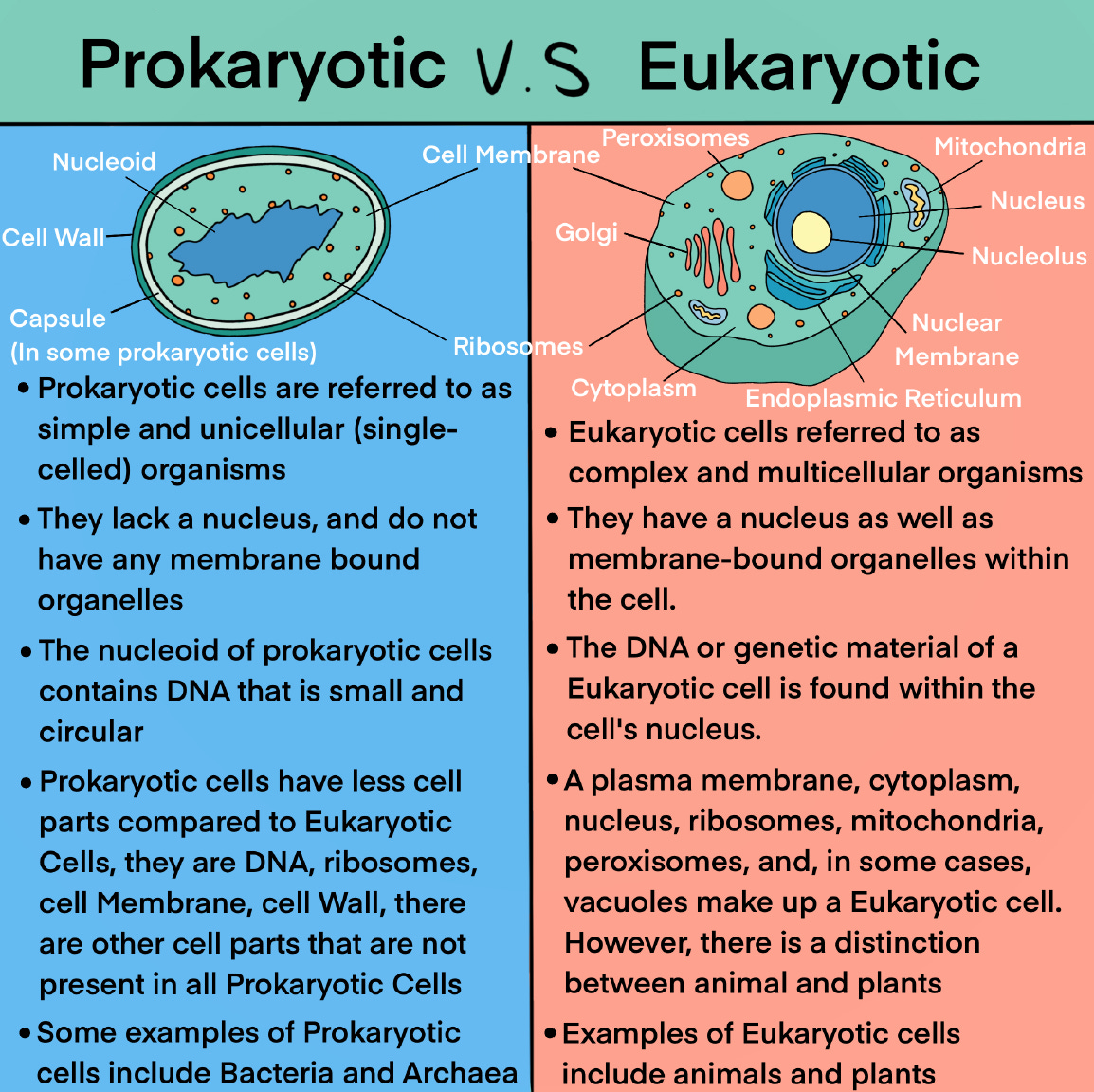
Bacteriophages, often referred to simply as phages, are viruses that infect and replicate within bacteria. They are the most abundant biological entities on Earth and play a crucial role in regulating bacterial populations, shaping microbial communities, and driving bacterial evolution. The process by which bacteriophages attack and enter bacteria varies depending on factors such as the type of bacteriophage and the structure of the bacterial cell wall, particularly whether the bacterium is Gram-positive or Gram-negative.
Bacteriophages will attack and enter, bacteria.
(Bifido is a gram positive bacteria. E. coli is gram negative—more on this in a moment.)
Mechanisms of Attack and Destruction by Phages: Gram Positive Bacteria versus Gram Negative:
1. Bacteriophage Attack and Entry into Gram-Positive Bacteria:
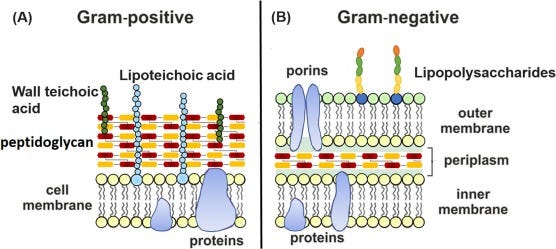
Gram-positive bacteria have a relatively simple cell wall structure consisting of a thick layer of peptidoglycan surrounded by a cytoplasmic membrane.
Attachment
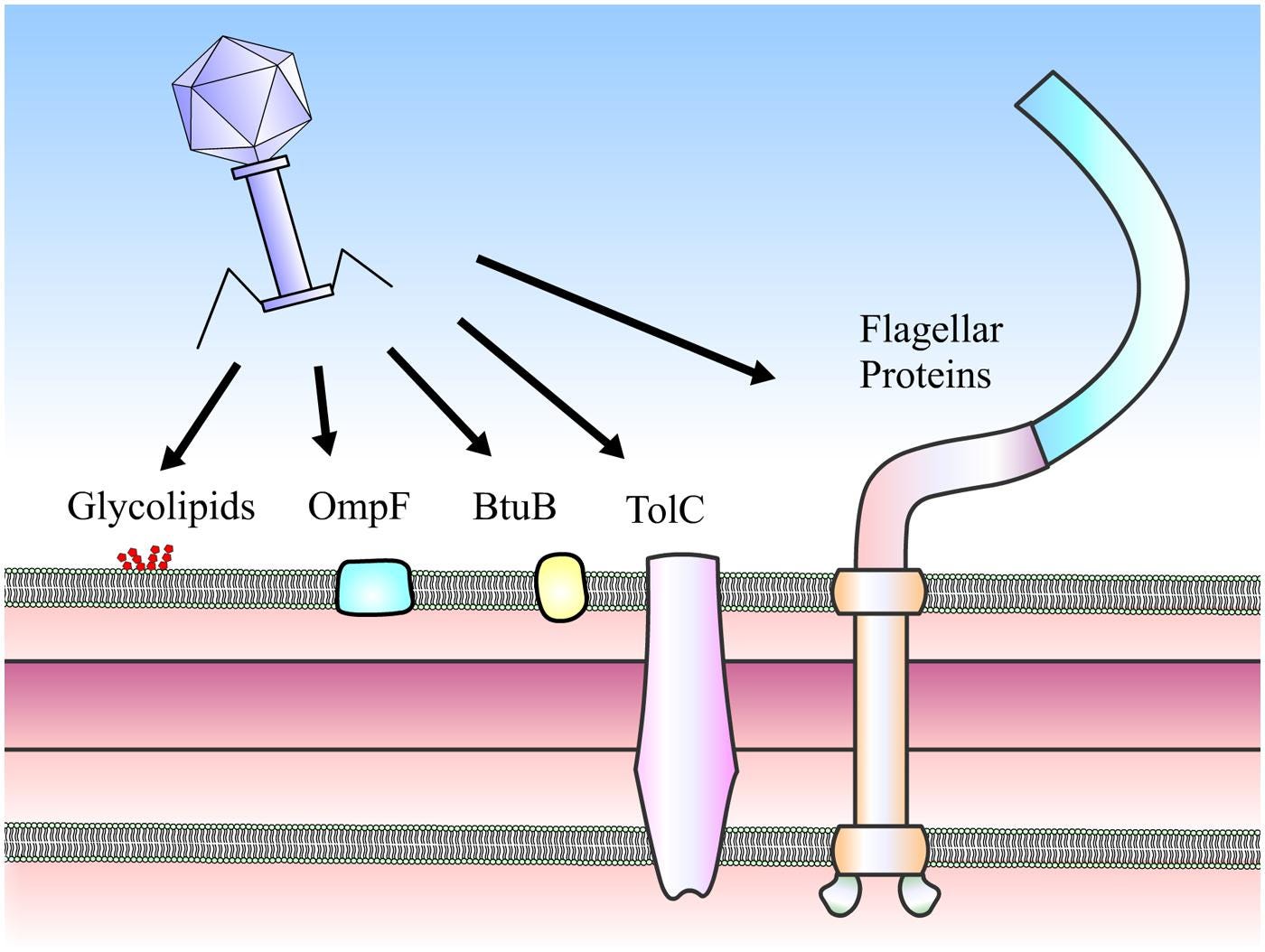
Bacteriophages have specific receptor-binding proteins on their surface that recognize and bind to specific receptors on the surface of the bacterial cell wall, like teichoic acids or other cell wall components.
The attachment is mediated by reversible interactions between viral and bacterial surface proteins.
Injection of Genetic Material
Once attached, the bacteriophage injects its genetic material (either DNA or RNA) into the bacterial cell through a specialized structure called a tail or tail fibers.
This injection process often involves the contraction of the tail sheath, which acts like a syringe to deliver the viral genetic material into the bacterial cytoplasm.
Replication and Assembly
Inside the bacterial cell, the viral genetic material hijacks the bacterial cellular machinery to replicate and produce new phage particles, involving the expression of phage genes that disrupt bacterial metabolism and redirect cellular resources towards phage replication.
New phage particles are assembled within the bacterial cytoplasm.
Lysis and Release:
Once assembled, the newly synthesized phage particles lyse (rupture) the bacterial cell wall, releasing progeny phages to infect neighboring bacteria.
This lysis is often mediated by the expression of phage-encoded lysins, enzymes that degrade the bacterial cell wall.
2. Bacteriophage Attack and Entry into Gram-Negative Bacteria:
Gram-negative bacteria have a more complex cell wall structure compared to Gram-positive bacteria, consisting of a thin layer of peptidoglycan sandwiched between an inner cytoplasmic membrane and an outer membrane. The process of bacteriophage infection in Gram-negative bacteria involves additional challenges due to the presence of the outer membrane. However, bacteriophages have evolved strategies to overcome these barriers:
Attachment and Adsorption
Bacteriophages first attach to specific receptors on the outer surface of the Gram-negative bacterial cell wall, like outer membrane proteins, lipopolysaccharides (LPS), or other cell surface molecules.
The attachment is often mediated by tail fibers or other viral surface proteins.
Penetration of the Outer Membrane
After attachment, some bacteriophages have specialized structures such as tail spikes or enzymes that facilitate the penetration of the outer membrane of Gram-negative bacteria.
These structures may degrade or disrupt components of the outer membrane, allowing the phage to access the underlying peptidoglycan layer.
Injection of Genetic Material
Once past the outer membrane, the bacteriophage injects its genetic material into the periplasmic space between the inner and outer membranes of the bacterial cell.
This injection may be facilitated by the contraction of the phage tail or other mechanisms.
Passage through the Inner Membrane
After injection into the periplasm, the phage genetic material must cross the inner cytoplasmic membrane to reach the bacterial cytoplasm by action of phage-encoded proteins that disrupt or penetrate the inner membrane.
Replication, Assembly, and Release
Once inside the bacterial cytoplasm, the phage genetic material hijacks the bacterial machinery to replicate and produce new phage particles.
The assembly and release of progeny phages may involve similar mechanisms as described for Gram-positive bacteria, ultimately leading to lysis of the host cell or, in some cases, the establishment of a lysogenic relationship where the phage genome integrates into the bacterial chromosome.
“Bacteriophages can be caudate, polyhedral, filamentous or pleomorphic (Figure 2) and, except for the caudate, they are not grouped into orders.”
We have what is called a gut VIROME.
The gut virome is incredibly diverse, comprising a vast array of bacteriophages that infect various bacterial species found in the gastrointestinal tract. While it's challenging to provide an exhaustive list of all bacteriophages present in the gut due to the immense diversity and ongoing discovery of new phages, here are some commonly studied families and genera of bacteriophages found in the gut:
Siphoviridae: This is one of the largest families of bacteriophages, characterized by long, non-contractile tails. Examples of Siphoviridae commonly found in the gut include:
Lactococcus phages: These infect bacteria belonging to the Lactococcus genus, commonly found in fermented dairy products and the gut microbiota of humans and animals.
Streptococcus phages: Streptococcus species are abundant in the oral cavity and the gut, and their phages have been extensively studied in the context of dental health and gastrointestinal infections.
Podoviridae: These bacteriophages have short, non-flexible tails and are known for their rapid adsorption and release cycles. Examples found in the gut include:
T7-like phages: T7 is a well-studied model phage used in molecular biology research. While not exclusive to the gut, T7-like phages have been identified in fecal samples and may play a role in gut microbiome dynamics.
Myoviridae: Bacteriophages in this family have contractile tails and are often characterized by their large size and complex structure. Examples in the gut include:
Escherichia coli phages: Escherichia coli is a common member of the gut microbiota, and its phages have been extensively studied due to their role in bacterial pathogenesis and their potential applications in phage therapy.
Caudovirales: This is an order of bacteriophages encompassing the families Siphoviridae, Podoviridae, and Myoviridae, among others. Caudovirales are abundant in diverse environments, including the gut.
Tailed phages: This is a broad category of bacteriophages characterized by their tail-like structures, which they use to attach to and infect bacterial cells. Tailed phages encompass various families and genera found in the gut and other environments.
Newsflash, COVID and the spike protein are not bacteriophages.
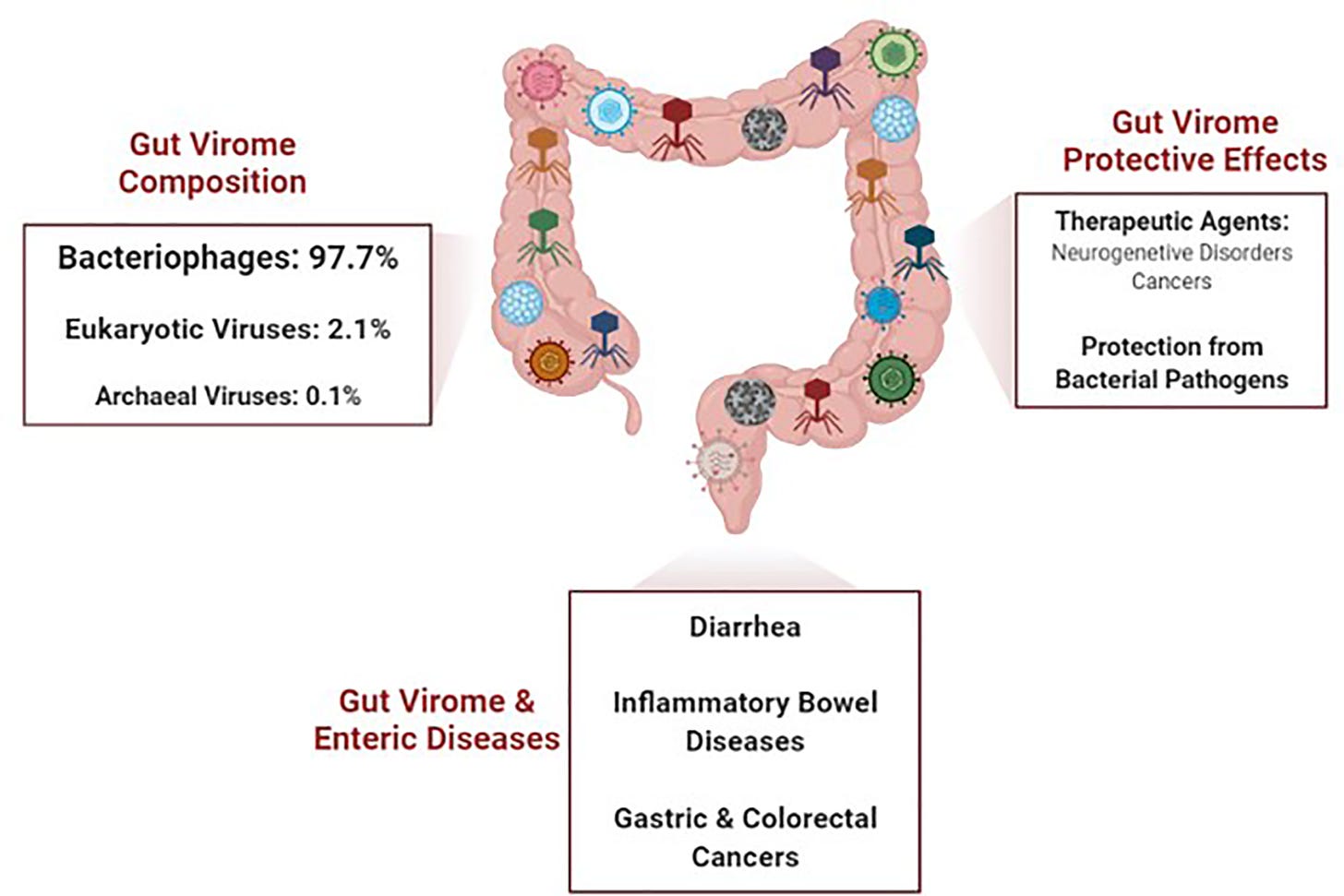
The virome in the gut refers to the collection of viruses inhabiting the gastrointestinal tract. It's a complex ecosystem comprising various viral species, primarily bacteriophages (viruses that infect bacteria), but also including eukaryotic viruses like adenoviruses and enteroviruses.
Bacteriophages: These are the most abundant components of the gut virome. They primarily infect bacteria, playing a significant role in modulating bacterial populations in the gut. Bacteriophages can influence the composition and diversity of the gut microbiome by selectively targeting certain bacterial species, not the human cells.
This process can indirectly impact host health by affecting microbial metabolism, immune responses, and gut homeostasis.Eukaryotic viruses: While less abundant than bacteriophages, eukaryotic viruses like adenoviruses and enteroviruses also inhabit the gut virome. These viruses can directly infect host cells in the gut lining, potentially causing gastrointestinal infections and inflammation—the HUMAN cells, not the bacteria.
Impact on the gut microbiome:
Bacterial population dynamics: Bacteriophages exert selective pressure on bacterial populations, leading to shifts in microbial composition and diversity. This can influence the stability and resilience of the gut microbiome.
Horizontal gene transfer: Bacteriophages can transfer genetic material between bacterial hosts through a process called transduction. This horizontal gene transfer can contribute to microbial evolution and adaptation within the gut ecosystem.
Microbial metabolism: Bacteriophages can alter microbial metabolism by lysing bacterial cells or modulating bacterial gene expression. This can affect nutrient utilization, fermentation pathways, and the production of metabolites such as short-chain fatty acids (SCFAs) that influence host health.
Immune modulation: Bacteriophages can interact with the host immune system, triggering immune responses that may impact gut inflammation, barrier function, and immune tolerance. The interplay between bacteriophages and the immune system is complex and can have both beneficial and detrimental effects on host health.
Imbalances in the gut virome composition have been linked to various gastrointestinal disorders, including inflammatory bowel disease (IBD), irritable bowel syndrome (IBS), and colorectal cancer.
Dysbiosis in the gut virome may contribute to disease pathogenesis through mechanisms such as increased inflammation, altered microbial metabolism, and impaired immune regulation.
Why COVID and other human respiratory viruses are NOT replication competent in bacterial cells compared to phages:
In the context of infecting bacterial cells, both viruses and bacteriophages can enter bacterial cells, but they do so through different mechanisms and have distinct outcomes:
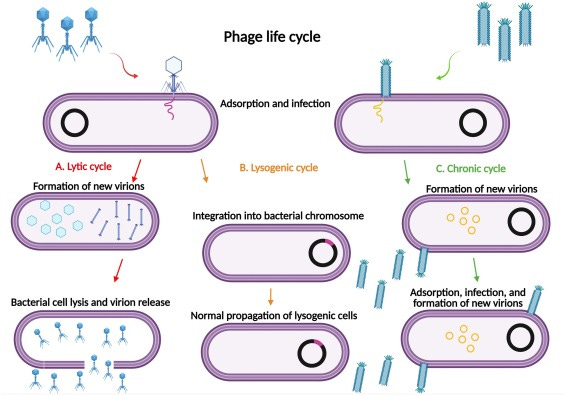
Bacteriophages
These are viruses that specifically infect bacteria. Bacteriophages have evolved to recognize and attach to specific receptors on the surface of bacterial cells. Once attached, they inject their genetic material (either DNA or RNA) into the bacterial cell. The phage genetic material then hijacks the bacterial cellular machinery, redirecting it to produce new phage particles. These new phage particles eventually lyse (burst) the bacterial cell, releasing more phages to infect neighboring bacteria. This process is known as the lytic cycle. However, some bacteriophages can also enter a dormant state within the bacterial cell, integrating their genetic material into the bacterial chromosome in a process called lysogeny. In this state, the phage is called a prophage and can replicate along with the bacterial cell's DNA until it is triggered to enter the lytic cycle.
Viruses (non-bacteriophages)
While viruses that infect bacteria are typically referred to as bacteriophages, it's worth noting that viruses that infect eukaryotic cells (e.g., animals, plants, fungi) can sometimes enter bacterial cells as well. However, this is not their primary mode of infection, and the outcomes of such interactions can vary. For example, some eukaryotic viruses may accidentally enter bacterial cells but fail to replicate due to incompatible cellular machinery. In other cases, they may be able to replicate to some extent but not produce viable viral particles. Eukaryotic viruses primarily target eukaryotic host cells. COVID is a eukaryotic virus. Bacteria are prokaryotic.
Therefore, the bacteria, especially Bifidobacterium, should be relatively safe from both COVID and the spike protein.
Next up, why GRAM NEGATIVE bacteria are used in biotech to make recombinant proteins, why a lipid nanoparticle is more likely to enter GRAM NEGATIVE negative bacteria (NOT GRAM POSITIVE BACTERIA LIKE BIFIDOBACTERIUM), house DNA biotech plasmids and more.