Thanks for hanging with me. Hopefully, some of you are connecting the dots now.
Let’s jump right in!
Lost microbes of COVID-19: Bifidobacterium, Faecalibacterium depletion and decreased microbiome diversity associated with SARS-CoV-2 infection severity
This study aimed to investigate the differences in gut microbiome diversity and composition between patients with varying severity of SARS-CoV-2 infection and PCR-negative exposed controls.
Study Design
The study utilized a cross-sectional design and conducted shotgun next-generation sequencing on stool samples obtained from patients with PCR-confirmed SARS-CoV-2 infections and PCR-negative exposed controls.
Patients were categorized based on the severity of their symptoms, ranging from asymptomatic to severe, following the National Institute of Health criteria. Exposed controls were individuals with prolonged or repeated close contact with SARS-CoV-2-infected patients or their samples, such as household members or frontline healthcare workers.
Microbiome Diversity and Composition Analysis
The researchers compared microbiome diversity and composition between patients and exposed controls at various taxonomic levels.
They examined factors such as the Shannon Index and Simpson Index to assess bacterial diversity.
Additionally, they analyzed the relative abundances of specific bacterial genera in patients versus controls.
Key Findings
Severely symptomatic SARS-CoV-2-infected patients exhibited significantly lower bacterial diversity compared to controls.
SARS-CoV-2-infected patients, regardless of symptom severity, had lower relative abundances of Bifidobacterium, Faecalibacterium, and Roseburium, while showing increased levels of Bacteroides.
Interestingly, there was an inverse association between disease severity and the abundance of these specific bacterial genera.
The dysbiosis pattern observed in the gut microbiome, characterized by decreased abundance of certain beneficial bacteria and increased abundance of Bacteroides, could serve as a marker for the severity of symptomatic SARS-CoV-2 infection. The authors suggest that interventions targeting the gut microbiome before, during, or after SARS-CoV-2 infection may help mitigate symptomatic severity.
Let’s be clear.—COVID did not cause the changes in the gut microbiome—they were already there.
How do we know that?
Let’s dive a bit further into details":
The study finds that SARS-CoV-2 positivity and infection severity are associated with decreased levels of protective bacteria, particularly Bifidobacterium and Faecalibacterium genera, as well as decreased bacterial diversity.
Bifidobacterium plays crucial immune functions, including enhancing Treg responses, reducing cell damage by inhibiting TNF-α and macrophages, and protecting against intestinal epithelial cell damage.
The abundance of Bifidobacterium decreases with age and obesity, both significant risk factors for SARS-CoV-2 infection.
Patients with more severe viral infections show decreased abundance of Bifidobacterium.
Faecalibacterium abundance is inversely related to SARS-CoV-2 positivity and infection severity.
Reduced levels of Faecalibacterium, particularly F. prausnitzii, are associated with aging, diabetes, obesity, and dietary factors, which are all risk factors for severe SARS-CoV-2 infection. T
he study suggests that Faecalibacterium levels may serve as indicators of overall human health and susceptibility to infection.
SARS-CoV-2 positivity and severity are also associated with decreased abundance of Roseburium and increased abundance of Bacteroides, although the implications of these changes remain unclear.
The study discusses the role of beneficial bacteria, such as Bifidobacterium and Faecalibacterium, in enhancing innate immunity and potentially modulating the inflammatory response associated with SARS-CoV-2 infection.
Reduced bacterial diversity and alterations in gut microbiota composition may contribute to the dysregulated immune response observed in severe cases of COVID-19.
Limitations and Future Directions
The study acknowledges limitations, including the lack of data on gut microbiome composition prior to SARS-CoV-2 infection onset.
So, no previous data available. NO smoking data. No drinking data. No antibiotic use data. Interesting.
I wonder what caused that?
Interesting.
Frequency, subtypes distribution, and risk factors of Blastocystis spp. in COVID-19 patients in Tehran, capital of Iran: A case-control study
”Results. The frequency of Blastocystis spp. in patients with COVID-19 (7.5%; 15/200 by molecular method vs. 6%; 12/200 by microscopy method) was slightly higher than in individuals without COVID-19 (4.5%; 9/200 by molecular method”
Next
Is co-infection of intestinal parasites with COVID-19 virus infection affecting its severity?
“High rate of co-infection with intestinal parasites was detected in group I. They were Blastocystis hominis (71.9%), Cryptosporidium spp., (65.9%), Entamoeba coli (63.0%), Cyclospora cayetanensis (54.1%), Isospora belli (51.1%), Entamoeba spp. (20.7%), and Giardia lamblia (8.9%) (Table 4).”
Let’s jump to the microbiome again and Covid:
Alterations in microbiota of patients with COVID-19: potential mechanisms and therapeutic interventions
post-acute COVID-19 syndrome=PACS
”Analyses of stool samples at admission disclosed that bacterial clusters distinctly differed between patients with and without PACS. Compared with the PACS patients, those without PACS-COVID-19 presented with gut bacterial compositions that were enriched for 19 bacteria and characterized by Bifidobacterium, Brautia, and Bacteroidetes. Patients with PACS displayed significantly lower gut bacterial diversity and richness than those of healthy controls. Thirteen bacterial species including Blautia wexlerae and Bifidobacterium longum were negatively associated with PACS at six months. Hence, these species may have protective roles during recovery from SARS-CoV-2 infection. In contrast, Actinomyces sp S6 Spd3, Actinomyces johnsonii, and Atopobium parvulum were positively correlated with PACS. The authors also reported that certain bacterial species such as Ruminococcus gnavus, Clostridium innocuum, and Erysipelatoclostridium ramosum remained variable from admission to the 6-month follow-up and were associated with several PACS symptoms.”
Feel free to do the additional reading on that image above, which is broken down in the linked study.
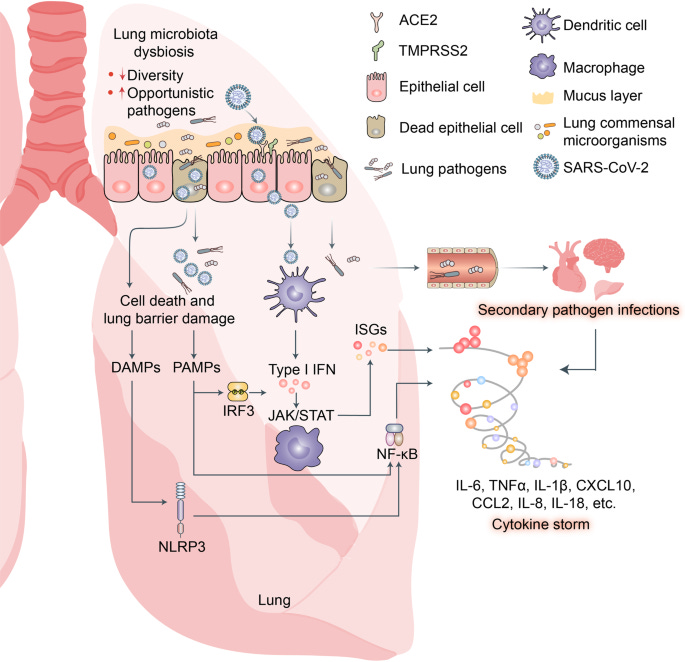
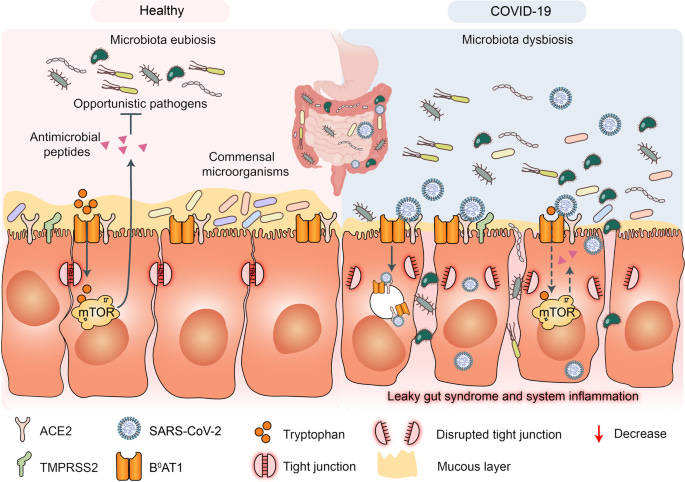
”Potential mechanisms of cytokine storm and secondary pathogen infections resulting from gut microbiota dysbiosis in patients with COVID-19. Gut microbiota are also disrupted by SARS-CoV-2 infection which potentially triggers cytokine storm and secondary pathogen infections. B0AT1 mediates neutral amino acid uptake by luminal surfaces of intestinal epithelial cells. It is also a molecular ACE2 chaperone. B0AT1 substrates such as tryptophan and glutamine activate antimicrobial peptide release, promote tight junction (TJ) formation, downregulate lymphoid proinflammatory cytokines, and modulate mucosal cell autophagy via mTOR signaling. As ACE2 is a molecular B0AT1 chaperone, ACE2-associated B0AT1 may be internalized during SARS-CoV-2 infection, decrease B0AT1 on cell membranes, promote gut opportunistic pathogen invasion, facilitate cytokine storms, and exacerbate COVID-19”
The important factor to note here, is that COVID DID NOT CAUSE the changes seen here. The LNP did not either.
Smoking
Drinking
Antibiotics
Parasites
Obesity
Increased Age
While these manifestations seem to occur in countries with fresh water, hygiene, and other concerns, COVID itself, The Spike Protein, and the LNP ain’t one of those.
There are CORRELATIONS.
(Next Stack will combine all of this in summary form)